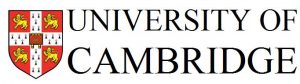[LEFT]: In epidermoid carcinoma cells, that the protein SON (green) is localising into nuclear speckles, a substructure in the nucleus.
[RIGHT]: SEPT9 (green) localizes to actin filaments in epidermoid carcinoma cells.
The first analysis of how proteins are arranged in a cell has been published today in Science, revealing that a large portion of human proteins can be found in more than one location in a given cell.
05.11.2017
Using the Sweden-based Cell Atlas, researchers examined the spatial distribution of the human proteome (the entire complement of proteins that make up the human body) that correspond to the majority of protein-coding genes. They described in unprecedented detail the distribution of proteins within the various substructures of the human body’s smallest unit, the cell.
Our cells contain ‘organelles’ – specialised substructures that carry out specific functions. These create partitions that form an enclosed environment for chemical reactions tailored to fulfill these functions. Since these functions are tightly linked to specific sets of proteins, knowing the subcellular location of the human proteome is key to understanding the function and underlying mechanisms of the human cell.
The study was led by Emma Lundberg, associate professor at KTH Royal Institute of Technology and responsible for the High Content Microscopy facility at the Science for Life Laboratory (SciLifeLab) in Stockholm, Sweden. The team generated more than 300,000 images to systematically resolve the spatial distribution of human proteins in cultivated cell lines, and map them to cellular compartments and substructures with single cell resolution.
The Cell Atlas is the result of more than 10 years of research within the Human Protein Atlas programme, and was launched in December 2016. The article in Science describes the detailed analysis of hundreds of thousands of images created as part of this international effort, which also involved groups in the China, South Korea, India, Denmark, and Germany.
“Only by studying the molecular components of the body’s smallest functional unit – the cell – can we reach a full understanding of human biology,” says KTH Professor Mathias Uhlen, director of the Human Protein Atlas.
The published article also includes a comparative study performed by Professor Kathryn Lilley, director of the Cambridge Centre for Proteomics, at Cambridge University, UK, which enabled the antibody-based immunofluorescence (IF) microscopy analysis to be validated by an alternative mapping strategy that used mass spectrometry, hyperLOPIT.
A total of 12,003 proteins targeted by 13,993 antibodies were classified into one or several of 30 cellular compartments and substructures, altogether defining the proteome of 13 major organelles. The organelles with the largest proteomes were the cytosol (4,279) and the nucleus (6,930) and its substructures, such as bodies and speckles.
Importantly, about one-half of the proteins are found in more than one compartment revealing a shared pool of proteins in functionally unrelated parts of the cell. This finding sheds new light on the complexity of cells.
”We have created the most detailed map of how proteins are arranged in a cell using two different high throughput approaches: high content imaging and spatial proteomics,” says Professor Lilley. ”Interestingly, we show a large proportion of human proteins can be found in more than one location in a given cell, overturning many pre-conceptions of how the cell operates.
”The Cell Atlas now provides us with new knowledge that will enable us to explore the functions of individual proteins and their role in human biology and disease.”
The Cell Atlas is an open access resource that can be used by researchers around the world to study proteins or organelles of interest. “The Atlas enables systems biology and cell modeling applications, and it is also a highly valuable resource for machine learning applications in image pattern recognition,” says Lundberg.
Reference
Thul, PJ et al. A subcellular map of the human proteome. Science; 11 May 2017; DOI: 10.1126/science.aal3321
Adapted from a press release from KTH Royal Institute of Technology in Stockholm.